Parameters and Presets
Parameters
Recipe parameters for Neurite Outgrowth and their descriptions are summarized in the table below.
Preset Group | Parameter Name | Min Value | Max Value | Description |
|---|---|---|---|---|
Detection | Background Removal Factor | 0 | 100 | Adjusts the sensitivity of the background removal operation; a lower value will preserve larger objects and more background variations |
Contrast Threshold | 0 | 255 (8-bit) 65,535 (16-bit) | Adjusts the detection sensitivity on the background-removed image; a lower value will detect bigger and more cell bodies | |
Fill Holes Size | 0 | 5,000 px2 or µm2 | Adjusts the maximum size threshold for filling in gaps inside a detected object; a lower value will preserve more holes in the detection | |
Smoothing Factor | 0 | 100 | Adjusts the amount of smoothing applied to the outlines of the detected objects; a lower value will preserve more of the objects' morphological features | |
Subset Filtering | Object Size | 0 | 1,000,000 px2 or µm2 | Specifies the range of objects to be included in the analysis results based on the areas of the detected objects |
Separation Factor (Cell Partition only) | 0 | 100 | Adjusts the sensitivity of the object separation operation; a lower value will preserve larger objects with multiple intensity peaks | |
Neurite Detection | Branch Sensitivity | 1 | 100 | Adjusts the sensitivity of the neurite tracing operation; a lower value will result in detection of fewer short branches |
Mean Neurite Width | 0.1 px or µm | 100 px or µm | Specifies the typical width of neurites on the image; a lower value will result in higher sensitivity for detection of thinner neurites | |
Min Neurite Length | 0 | 1,000 px or µm | Specifies the minimum length of neurite branches to be included in the analysis result |
Presets
There are three preset groups in the recipe: Detection, Subset Filtering and Neurite Detection. Each group has three pre-configured parameter groupings to help you get started on the analysis. The default preset values are as follows:
Results
Measurements
The Neurite Outgrowth recipe generates morphological, intensity, and count measurements for each detected neuron, including the total number of neurites per cell. You can add additional measurements to the analysis results with the Measurement Tool in Aivia and view measurement definitions on the Measurement Definitions page. The measurements generated by the recipe are given in the table below.
Object Set | Morphology | Intensity | Count | Advanced |
|---|---|---|---|---|
Soma |
|
|
|
|
Neurite | None | None |
| None |
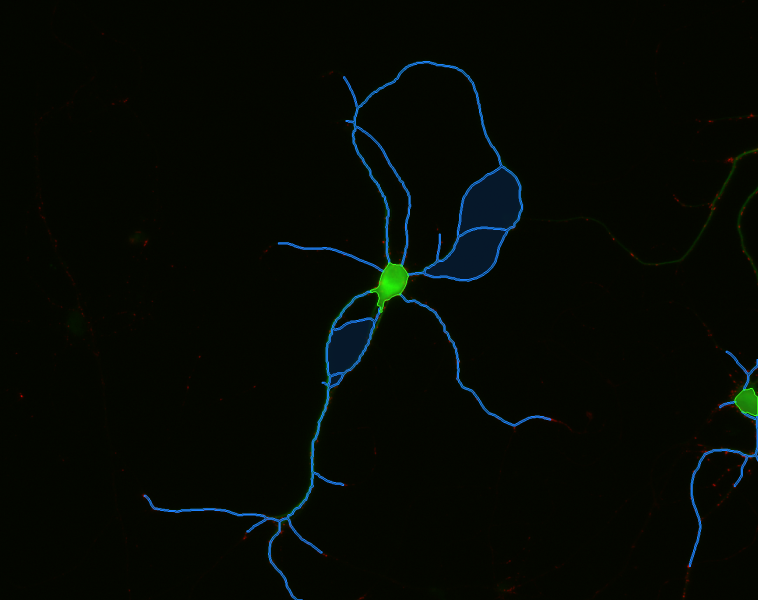
Image Credits
Ginger Withers, Whitman College, Cell Image Library (CIL:12566), http://www.cellimagelibrary.org/images/12566
Related Articles
Related articles appear here based on the labels you select. Click to edit the macro and add or change labels.
|