Automatic Parameter Selection
Parameters and Presets
Parameters
Recipe parameters for 3D Neuron Analysis - FL and their descriptions are summarized in the table below.
Preset Group | Parameter Name | Min Value | Max Value | Description |
|---|---|---|---|---|
Soma Detection | Soma Diameter | 0 | 1,000 (px or µm) | Specifies the average diameter of somas (neuron cell bodies); a lower value leads to detection of smaller objects on the image |
Min Intensity Threshold (Enable Soma Partition only) | 0 | 255 (8-bit) 65,535 (16-bit) | Specifies the minimum intensity of somas on the image for partitioning touching somas; this parameter is only enabled when the Enable Soma Partition option is selected; a lower value reduces the number of partitions and preserves larger soma | |
Dendrite Detection | Dendrite Diameter | 0 | 50 (px or µm) | Specifies the diameter of a typical dendrite on the image; a lower value increases detection sensitivity for thin dendrites and decreases detection sensitivity for thicker dendrites (such as axons) |
Min Branch Length | 0 | 1,000 (px or µm) | Specifies the minimum length of a traced branch to be included in the analysis results | |
Intensity Threshold | 0 | 255 (8-bit) 65,535 (16-bit) | Specifies the intensity range for dendrite tracing using the voxel scooping algorithm; the lower threshold specifies the minimum intensity that is not considered background; the upper threshold specifies the minimum intensity above which the voxels are always considered as part of a dendrite | |
Mean Intensity Offset | 0 | 255 (8-bit) 65,535 (16-bit) | Adjusts the detection sensitivity of the dendrite tracing algorithm; a higher value reduces the number of dendrites detected that are at or near the minimum intensity threshold specified above | |
Max Connection Distance (Skip Soma Detection only) | 0 | 1,000 (px or µm) | Specifies the maximum gap distance between dendritic traces to join together; this parameter is only enabled when the Skip Soma Detection option is selected; a lower value increases the number of disconnected dendritic branches | |
Spine Detection (Enable Spine Detection and Enable Enhanced Spine Detection only) | Typical Spine Head Diameter | 0 | 100 (px or µm) | Specifies the approximate diameter of a typical dendritic spine head; this parameter is only enabled when the Enable Spine Detection option or Enable Enhanced Spine Detection option is selected |
Spine Head Diameter | 0 | 100 (px or µm) | Specifies the size range for a spine to be included in the analysis results based on the diameter of the spine head; this parameter is only enabled when the Enable Spine Detection option or Enable Enhanced Spine Detection option is selected | |
Max Spine Neck Length | 0 | 100 (px or µm) | Specifies the maximum distance between spine heads and dendritic branches; this parameter is only enabled when the Enable Spine Detection option or Enable Enhanced Spine Detection option is selected; a lower value reduces the search range for spine heads |
Presets
There are three preset groups in the recipe: Soma Detection, Dendrite Detection, and Spine Detection; each group has three pre-configured parameter groupings to help you get started on the analysis. The default preset values are given in the sections to follow.
Measurements
The 3D Neuron Analysis - FL recipe generates morphological, intensity, and count measurements for the somas, dendrites, and spines of each detected neuron. You can add additional measurements to the analysis results by using the Measurement Tool in Aivia and view measurement definitions on the Measurement Definitions page. The measurements generated by the recipe are summarized in the table below.
Object Set | Morphology | Intensity | Count |
|---|---|---|---|
Soma Set |
|
| None |
Dendrite Trees |
| None |
|
Dendrite Segments |
|
|
|
Spines |
| None | None |
Spine Heads |
|
| None |
Spine Necks |
|
| None |
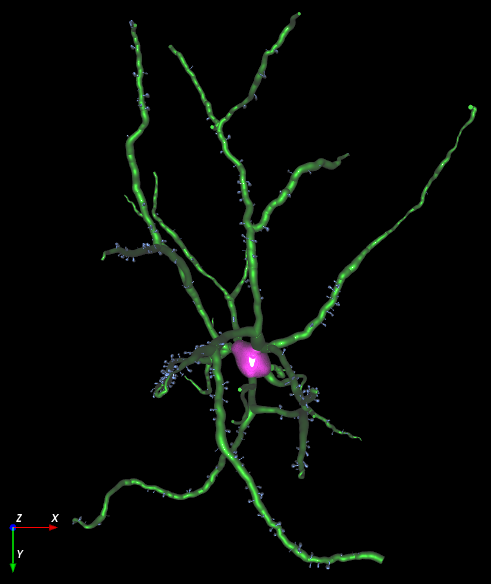
Image Credits
Maryann Martone and Eric Bushong, Cell Image Library (CIL:48401), http://www.cellimagelibrary.org/images/48401
Related Articles
Related articles appear here based on the labels you select. Click to edit the macro and add or change labels.
|